Luigi Warren
New Allele Biotech Publication on Stem Cells
Feeder-Free Reprogramming of Human Fibroblasts with Messenger RNA
Current Protocols in Stem Cell Biology • November 13, 2013
DOI: 10.1002/9780470151808.sc04a06s27
Authors: Luigi Warren, Jiwu Wang
This unit describes a feeder-free protocol for deriving induced pluripotent stem cells (iPSCs) from human fibroblasts by transfection of synthetic mRNA. The reprogramming of somatic cells requires transient expression of a set of transcription factors that collectively activate an endogenous gene regulatory network specifying the pluripotent phenotype. The necessary ectopic factor expression was first effected using retroviruses; however, as viral integration into the genome is problematic for cell therapy applications, the use of footprint-free vectors such as mRNA is increasingly preferred. Strong points of the mRNA approach include high efficiency, rapid kinetics, and obviation of a clean-up phase to purge the vector. Still, the method is relatively laborious and has, up to now, involved the use of feeder cells, which brings drawbacks including poor applicability to clinically oriented iPSC derivation. Using the methods described here, mRNA reprogramming can be performed without feeders at much-reduced labor and material costs relative to established protocols.
Allele iPSC Service and Technology Licensing Contact: http://www.allelebiotech.com/cell-line-and-culture-services/#ips-line
New Allele Product of the Month: FP-nAb™ products for 100% pull-down
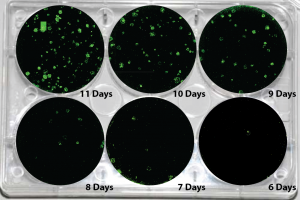
Picture Blog — Human mRNA-Induced Pluripotent Stem Cells Generated in Days
R-iPS Cell FAQ 1:
What phenotypic changes can be observed during a successful reprogramming trial?
About one week out, target fibroblasts should show an involution of fibroblastic processes, and foci or clusters of epitheliod cells—ideally with small nuclei, minimal cytoplasm, and signs of ongoing mitosis—should appear. Colonies with hESC morphology typically start emerging in ~10-14 days.
Methods of iPSC Generation Update
Induced pluripotent stem cells can be directly generated from adult cell cultures through the introduction of a group of factors, e.g. Oct4, Sox2, Klf4, and c-Myc (the Yamanaka factors) [Takahashi and Yamanaka, 2006]. Additional factors such as Nanog and Lin28 can either substitute some of the Yamanaka factors or supplement them for higher reprogramming efficiency [Yu et al. 2007].
The original pluripotent stem cells induction methods involved retrovirus or lentivirus that would leave foot-print in the host genome, a concern for clinical use of iPSCs. Several groups have tried to create iPSCs without integrating viruses, such as using small molecules, directly delivering proteins instead of cDNAs, viruses with RNA genomes, episomal systems, or removable elements such as PiggyBac or Sleeping Beauty transposons. From the literature and our first-hand experience in the iPS market, none of these methods has become a widely applicable tool, mostly due to impractically low reprogramming efficiency.
In addition to low efficiency, RNA viruses, such as the sendai virus, are still viruses and have virus-associated risks. Episomal plasmids or removable transposons still involve DNA, so the possibility of genomic integration by recombination remains. In case of some transposons such as PiggyBac, there is an additional question about the degree of removal – whether it is certain that all integrated transposons, often inserted within genes, are deleted; in case of transposons similar to Sleeping Beauty, the small footprints they leave behind may post a concern.
The method of choice for generating zero-footprint iPSCs should clearly be RNA-based without the involvement of virus. Luigi Warren and his former colleagues at Harvard demonstrated that by using in vitro transcribed iPS factor mRNAs with modified CTP and UTP, and 5’-cap can effectively reprogram a number of different human as well as mouse cells. The efficiency even exceeds those by using retrovirus or lentivirus by 10 to 100 fold. Furthermore, the RiPSCs created with mRNAs appear to be closer to hESCs as shown by expression profiling.
Very recently, a few miRNAs that have high expression levels in stem cells were shown to be able to reprogram mouse and human somatic cells when expressed together from a lentivirus [Anokye-Danso et al. 2011]. while that work used lentivirus, thus not directly applicable to the current project, Miyoshi et al. later showed that by using synthesized mature miRNA (overlapping but not the same set of miRNAs as used by Anokye-Danso et al.) reprogramming cold be achieved without viral infection. We believe that this is a promising method and would like to pursue it further and to find out whether these mi-iPSCs relate to hESCs as closely as R-iPSCs. Because transfecting synthetic miRNAs “does operate at considerably lower efficiency” in terms of iPSC creation [Miyoshi et al. 2011],alternative protocols may include transfecting the iPS factor mRNAs together with various miRNAs at different doses and frequencies.
New Product of the Week: EF1a-lacZ lentivirus particles, for expressing nuclear lacZ in virtually any human or mouse cells.
This week save 25% on photoconvertible fluorescent protein mClavGR2 cloning plasmids. Email oligo@allelebiotech.com with code FPBLOG0831.
Blog References: Warren, L. et al. “Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA” 2010 Cell Stem Cell 7(5): 618-30
Anokye-Danso, F. et al. “Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency” 2011 Cell Stem Cell 8(4): 376-88
Miyoshi, N. et al. “Reprogramming of mouse and human cells to pluripotency using mature microRNAs” 2011 Cell Stem Cell 8(6): 633-8
Kim, H. et al. “miR-371-3 expression predicts neural differentiation propensity in human pluripotent stem cells” 2011 Cell Stem Cell 8(6): 695-706
Takahashi, K. and Yamanaka, S. “Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors” 2006 Cell 126(4): 663-76
Yu, J. et al. “Induced pluripotent stem cell lines derived from human somatic cells” 2007 Science 318(5858): 1917-20
Categories
- Allele Mail Bag
- cGMP
- Customer Feedback
- Fluorescent proteins
- iPSCs and other stem cells
- nAb: Camelid Antibodies, Nanobodies, VHH
- Next Generation Sequencing (NextGen Seq)
- NIH Budget and You
- oligos and cloning
- Open Forum
- RNAi patent landscape
- SBIR and Business issues
- State of Research
- Synthetic biology
- Uncategorized
- Viruses and cells
- You have the power
Archives
- October 2018
- April 2018
- March 2018
- January 2018
- October 2017
- September 2017
- August 2017
- March 2017
- February 2017
- January 2017
- November 2016
- September 2016
- August 2016
- July 2016
- June 2016
- May 2016
- April 2016
- February 2016
- October 2015
- September 2015
- August 2015
- June 2015
- March 2015
- January 2015
- December 2014
- March 2014
- February 2014
- January 2014
- December 2013
- November 2013
- October 2013
- September 2013
- August 2013
- July 2013
- June 2013
- May 2013
- April 2013
- March 2013
- January 2013
- December 2012
- November 2012
- October 2012
- September 2012
- August 2012
- July 2012
- May 2012
- April 2012
- February 2012
- January 2012
- December 2011
- November 2011
- October 2011
- September 2011
- August 2011
- July 2011
- June 2011
- May 2011
- April 2011
- March 2011
- February 2011
- January 2011
- December 2010
- November 2010
- October 2010
- September 2010
- August 2010
- July 2010
- June 2010
- May 2010
- April 2010
- March 2010
- February 2010
- January 2010
- December 2009
- November 2009
- October 2009
- September 2009
- August 2009
- July 2009
- June 2009
- May 2009
- April 2009
- March 2009
- February 2009
- January 2009
- December 2008
- October 2008
- August 2008
- July 2008